This afternoon, I got reply from Stephanie Nicole Lewis from Virginia Polytechnic Institute and State University. 
Maybe I can choose water as flexible residue for better docking result. The protein and water all be treated as the receptor for the following docking. I use Autodock to open total.pdb, then add the polar hydrogen atoms. Then I used VMD to extract protein and glycan of the protein crystal PDB file and save to another PDB file. I used VMD to extract water and add hydrogen atoms by chimera and save to PDB. Although I am not sure whether the hydrogen atoms are optimized or not, at least they did not orient to one direction.Īutodock seems have trouble to deal with the structure got from chimera. And finally, I found USCF chimera has a module to add hydrogen atoms very well. I tried some other programs, VMD, Schrodinger, HAAD server, WHAT IF. So I just do that for the job now.Īutodock can not add hydrogen atoms on the lonely oxygen atoms in x-ray crystal structure. 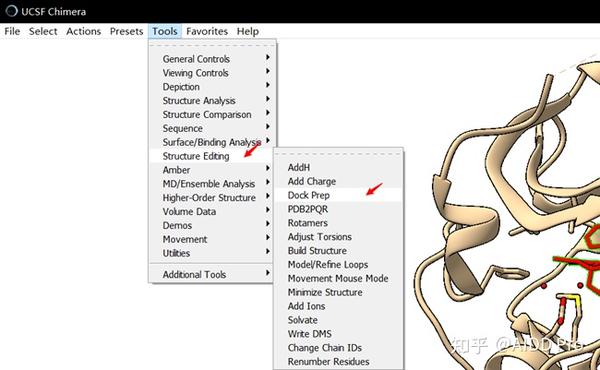
By searching for a while, it seemed the docking with water only use the water inside the crystal structure. I am confused whether I should add water manually or just use the crystal water. I was asked to do docking with explicit water.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
